The major take-aways of this research include the following: (1) DNA sequencing can be used to identify residual FLT3-ITD or NPM1 variants in the blood of adults with AML who are in their first remission before allogeneic HSCT; (2) The persistence of FLT3-ITD or NPM1 variants in pretransplant blood samples was associated with an increased post-HSCT relapse rate; (3) Patients with persistent variants in pretransplant blood samples had significantly lower post-HSCT survival rates than those who did not.
Persistent FLT3 internal tandem duplication (ITD) or NPM1 variants in pretransplant blood samples from patients with acute myeloid leukemia (AML) who were in their first remission were associated with “increased relapse and worse survival,” according to a recent study.
Laura Dillon, PhD, of the National Heart, Lung, and Blood Institute at the National Institutes of Health, and colleagues conducted the Pre-MEASURE study because there is “currently no standard” for testing measurable residual disease (MRD) in AML.
For example, flow cytometry is “commonly used” for MRD testing in AML, but “concerns have been raised about lack of interlaboratory standardization, leading to potentially limited prognostic value in decentralized settings,” Dr. Dillon and colleagues wrote.
Due to the lack of a standard testing method, Dr. Dillon and colleagues used targeted deep DNA sequencing to test the hypothesis that detecting “specific residual AML-associated variants” in the blood of patients who are in their first remission prior to allogeneic hematopoietic stem cell transplantation (HSCT) would be “associated with higher rates of relapse and mortality after transplant.”
Retrospective Study Tests Pretransplant Blood Samples From Patients with AML
The retrospective, observational study included adults who received their first allogeneic HSCT during their first remission between 2013 and 2019. Dr. Dillon and colleagues performed targeted DNA sequencing on banked pretransplant blood samples from the patients, who had AML associated with variants in FLT3, NPM1, IDH1, IDH2, or KIT. They defined MRD-positive results as those showing a variant allele fraction of 0.01% or higher.
The researchers analyzed 1,075 patients, identifying 822 patients who had FLT3-ITD, mutated NPM1, or both. The median patient age was 57.1 years, 54% of patients were female, and 84% were White. The researchers grouped patients into a discovery cohort that included those who received a transplant between 2013 and 2017 (n=371) and a validation cohort that included those who received a transplant between 2018 and 2019 (n=451).
The gene combinations used for next-generation sequencing (NGS) were determined by the discovery cohort and validated in the validation cohort. The study’s primary outcomes were overall survival (OS) and relapse rates.
Persistent Variants Before Transplant Linked with Post-HSCT Relapse, Survival Outcomes
In the discovery cohort, pretransplant remission blood samples from 64 patients (17.3%) showed persistent FLT3-ITD, mutated NPM1, or both.
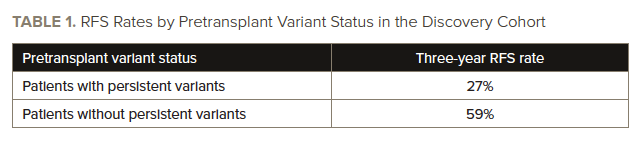
The three-year relapse rate was 59% in patients with persistent variants, significantly higher than the rate of 24% in those who did not have persistent variants (hazard ratio [HR], 3.71; 95% CI, 2.55-5.41; P<.001). The three-year OS rate was significantly lower in patients with persistent variants (34%) than in those without persistent variants (66%; HR, 2.60; 95% CI, 1.85-3.65; P<.001). See TABLE 1 for information on three-year relapse-free survival (RFS) rates in the discovery cohort.
In the validation cohort, pretransplant remission blood samples from 78 patients (17.3%) showed persistent FLT3-ITD, mutated NPM1, or both.
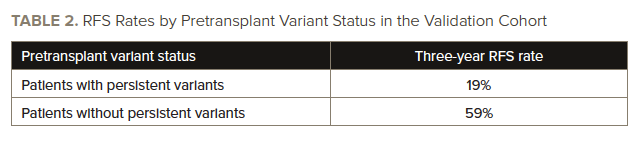
Patients with persistent variants had a three-year relapse rate of 68%, significantly higher than the rate of 21% in those who did not have persistent variants (HR, 4.32; 95% CI, 2.98-6.26; P<.001). Patients with persistent variants had a three-year survival rate of 39%, significantly lower than the rate of 63% in patients who did not have persistent variants (HR, 2.43; 95% CI, 1.71-3.45; P<.001). See TABLE 2 for information on three-year RFS rates in the validation cohort.

While persistent variants were associated with “higher rates of relapse and worse survival,” this issue was “partially mitigated” in younger patients who received high-intensity myeloablative conditioning (three-year relapse rate, 53% vs 78%; HR, 1.97; 95% CI, 1.03-3.75; P=.04), according to Dr. Dillon and colleagues.
Identifying Differential Risk
The authors outlined several limitations of the study, noting that it is “unknown” how results from bone marrow samples would differ from the results they found in blood samples.
Dr. Dillon and colleagues also did not have access to pretransplant flow cytometry MRD testing data from patients and could not evaluate if the NGS results would be consistent with those results. They noted that it is “unknown how these results apply to others who did not undergo transplant” and only around 10% of patients studied received maintenance therapy.
However, despite the study’s limitations, its results showed NGS MRD testing on pretransplant blood samples from the first remission in patients who have AML with FLT3-ITD, NPM1 mutations, or both “could identify differential risk between individuals otherwise placed in the same baseline risk classification.”
“Among patients with [AML] in first remission prior to allogeneic hematopoietic cell transplant, the persistence of FLT3 internal tandem duplication or NPM1 variants in the blood at an allele fraction of 0.01% or higher was associated with increased relapse and worse survival compared with those without these variants,” Dr. Dillon and colleagues concluded. “Further study is needed to determine whether routine DNA-sequencing testing for residual variants can improve outcomes for patients with [AML].”
Reference
Dillon LW, Gui G, Page KM, et al. DNA sequencing to detect residual disease in adults with acute myeloid leukemia prior to hematopoietic cell transplant. JAMA. 2023;329(9):745-755. doi:10.1001/jama.2023.1363




 © 2025 Mashup Media, LLC, a Formedics Property. All Rights Reserved.
© 2025 Mashup Media, LLC, a Formedics Property. All Rights Reserved.